Transcriptomics of Pseudomonas aeruginosa PA14 upon deletion of the sigma factor RpoS
AUTHORS
Luis Mauricio Salazar-Garcia 1, Jose Manuel Villalobos-Escobedo 2
Industrial Genomics Laboratory, FEMSA Biotechnology Center, School of Engineering and Sciences, Tecnológico de Monterrey, Monterrey, Nuevo Leon, Mexico.1
Integrative Biology Research Unit, The Institute for Obesity Research, Tecnológico de Monterrey, Monterrey, Nuevo Leon, Mexico.2
e-mail: jose.villalobos@tec.mx
DOWNLOAD THE PAPER
Understanding bacterial resilience through genomics
This study deciphers how Pseudomonas aeruginosa—one of the world’s most persistent opportunistic pathogens—reorganizes its gene expression when it loses its master regulator: the sigma factor RpoS (σ^S).
Using CRISPR/Cas9 genome editing, researchers generated a mutant of the virulent PA14 strain by introducing a STOP codon into the rpoS gene, completely deactivating its function. This modification allowed them to observe how the bacterium responds at the molecular level when its stress-defense system is turned off.
Loss of RpoS triggers a massive reprogramming of the transcriptome, altering metabolic pathways that sustain the bacterium’s resistance and infectious capacity.
The sigma factor RpoS not only regulates survival under adverse conditions such as nutrient limitation or oxidative stress but also influences the expression of virulence genes.
This study provides a detailed view of the RpoS regulon in the highly virulent PA14 strain—offering new insights into the mechanisms that make P. aeruginosa such a successful organism in both clinical and industrial environments.
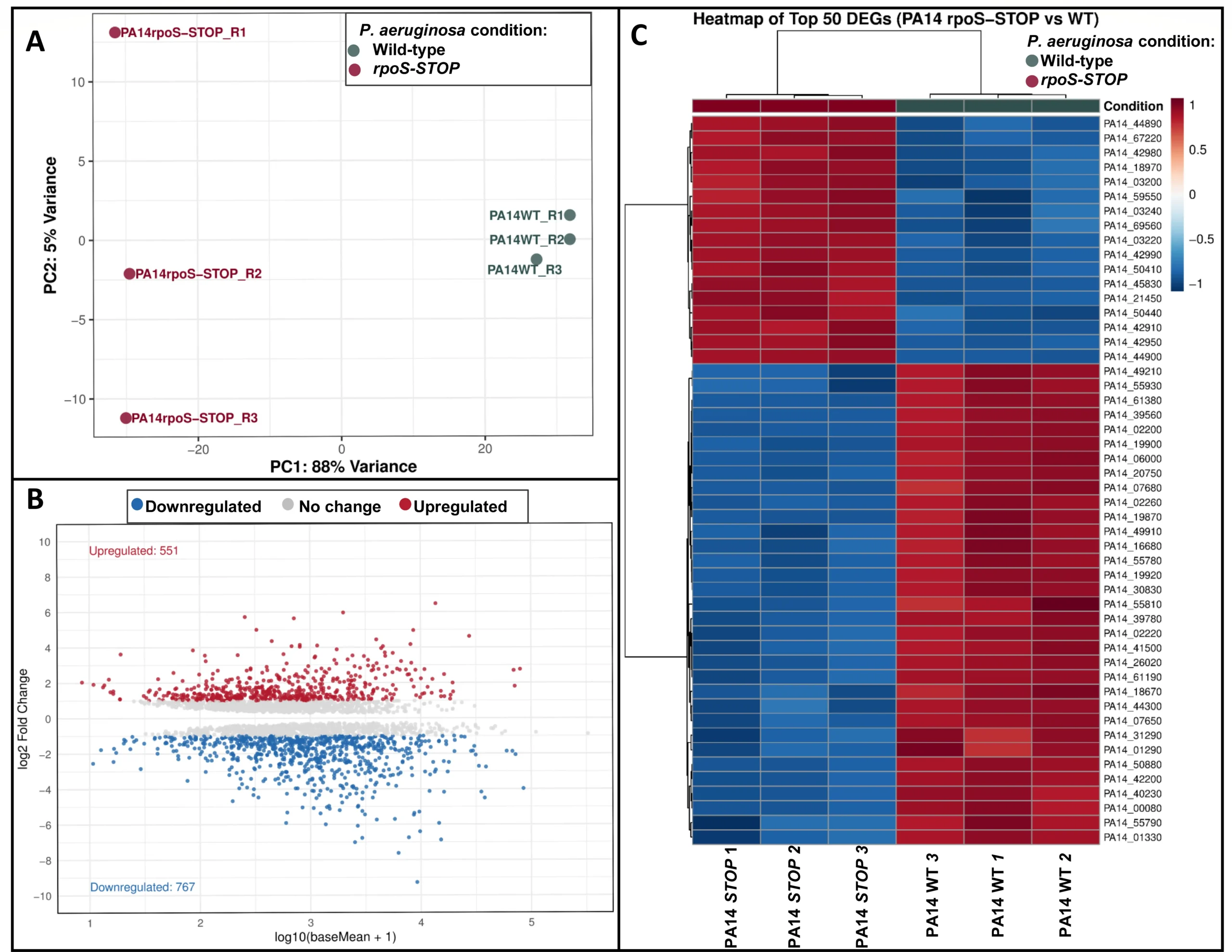
Fig 1: Análisis transcriptómico de cepas de tipo salvaje PA14 de P. aeruginosa y cepas mutantes rpoS-STOP .
- Lyczak JB, Cannon CL, Pier GB. 2000. Establishment of Pseudomonas aeruginosa infection: lessons from a versatile opportunist. Microbes Infect 2:1051–1060.
- Estahbanati HK, Kashani PP, Ghanaatpisheh F. 2002. Frequency of Pseudomonas aeruginosa serotypes in burn wound infections and their resistance to antibiotics. Burns 28:340–348.
- Sadikot RT, Blackwell TS, Christman JW, Prince AS. 2005. Pathogen–host interactions in Pseudomonas aeruginosa pneumonia. Am J Respir Crit Care Med 171:1209–1223.
- Diggle SP, Whiteley M. 2020. Microbe Profile: Pseudomonas aeruginosa: opportunistic pathogen and lab rat. Microbiology (Reading) 166:30–33.
- Denissen J, Reyneke B, Waso-Reyneke M, Havenga B, Barnard T, Khan S, Khan W. 2022. Prevalence of ESKAPE pathogens in the environment: antibiotic resistance status, community-acquired infection and risk to human health. Int J Hyg Environ Health 244:114006.
- Lee DG, Urbach JM, Wu G, Liberati NT, Feinbaum RL, Miyata S, Diggins LT, He J, Saucier M, Déziel E, Friedman L, Li L, Grills G, Montgomery K, Kucherlapati R, Rahme LG, Ausubel FM. 2006. Genomic analysis reveals that Pseudomonas aeruginosa virulence is combinatorial. Genome Biol 7:1–14.
- Wolfgang MC, Kulasekara BR, Liang X, Boyd D, Wu K, Yang Q, Miyada CG, Lory S. 2003. Conservation of genome content and virulence determinants among clinical and environmental isolates of Pseudomonas aeruginosa. Proc Natl Acad Sci USA 100:8484–8489.
- Suh SJ, Silo-Suh L, Woods DE, Hassett DJ, West SE, Ohman DE. 1999. Effect of rpoS mutation on the stress response and expression of virulence factors in Pseudomonas aeruginosa. J Bacteriol 181:3890–3897.
- Whiteley M, Parsek MR, Greenberg EP. 2000. Regulation of quorum sensing by RpoS in Pseudomonas aeruginosa. J Bacteriol 182:4356–4360.
- Schuster M, Lostroh CP, Ogi T, Greenberg EP. 2003. Identification, timing, and signal specificity of Pseudomonas aeruginosa quorum-controlled genes: a transcriptome analysis. J Bacteriol 185:2066–2079.
- Muriel-Millán LF, Montelongo-Martínez LF, González-Valdez A, Bedoya-Pérez LP, Cocotl-Yañez M, Soberón-Chávez G. 2024. The alternative sigma factor RpoS regulates Pseudomonas aeruginosa quorum sensing response by repressing the pqsABCDE operon and activating vfr. Mol Microbiol 121:291–303.
- Mathee K, Narasimhan G, Valdes C, Qiu X, Matewish JM, Koehrsen M, Rokas A, Yandava CN, Engels R, Zeng E, Olavarietta R, Doud M, Smith RS, Montgomery P, White JR, Godfrey PA, Kodira C, Birren B, Galagan JE, Lory S. 2008. Dynamics of Pseudomonas aeruginosa genome evolution. Proc Natl Acad Sci USA 105:3100–3105.
- Freschi L, Vincent AT, Jeukens J, Emond-Rheault J-G, Kukavica-Ibrulj I, Dupont M-J, Charette SJ, Boyle B, Levesque RC. 2019. The Pseudomonas aeruginosa pan-genome provides new insights on its population structure, horizontal gene transfer, and pathogenicity. Genome Biol Evol 11:109–120.
- Volke DC, Martino RA, Kozaeva E, Smania AM, Nikel PI. 2022. Modular (de)construction of complex bacterial phenotypes by CRISPR/nCas9-assisted, multiplex cytidine base-editing. Nat Commun 13:3026.
- Chen S, Zhou Y, Chen Y, Gu J. 2018. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890.
- Kim D, Paggi JM, Park C, Bennett C, Salzberg SL. 2019. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol 37:907–915.
- Liao Y, Smyth GK, Shi W. 2014. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30:923–930.
- Love MI, Huber W, Anders S. 2014. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550.



