Genomics and metabolomics of Trichoderma harzianum T9: a desert fungus with potential for sustainable agriculture
AUTHORS
Francisco Vargas-Gasca1, Enrique Pola-Sánchez2, Ana Valeria García-Lartigue2, Alan D. Gomez-Vargas3, Pablo Cruz-Morales4, Ana Calheiros Carvalho4, Daniela Rago4, Linda Ahonen4, Elva Teresa Aréchiga-Carvajal5, José Manuel Villalobos-Escobedo2, Vianey Olmedo-Monfil1
Universidad de Guanajuato.1
Institute for Obesity Research.2
CINVESTAV Irapuato.3
Technical University of Denmark.4
Universidad Autónoma de Nuevo León.5
e-mail: jose.villalobos@tec.mx
download the paper
The use of microorganisms as alternatives to chemical pesticides has become a priority in achieving more sustainable agriculture.
Facing the challenges of climate change and soil degradation, we sequenced the genome of Trichoderma harzianum T9 — a strain isolated from the arid, alkaline soils of Mina, Nuevo León — capable of thriving where almost nothing grows.
This desert fungus not only withstands extreme temperatures and high-pH soils; it also inhibits highly aggressive phytopathogens such as Macrophomina spp. and Neopestalotiopsis rosae, which cause significant economic losses in Mexico’s strawberry crops.
The genome of T. harzianum T9 represents a step forward toward more resilient and sustainable agriculture — a biotechnology emerging from the Mexican desert that could transform the way we protect our crops.

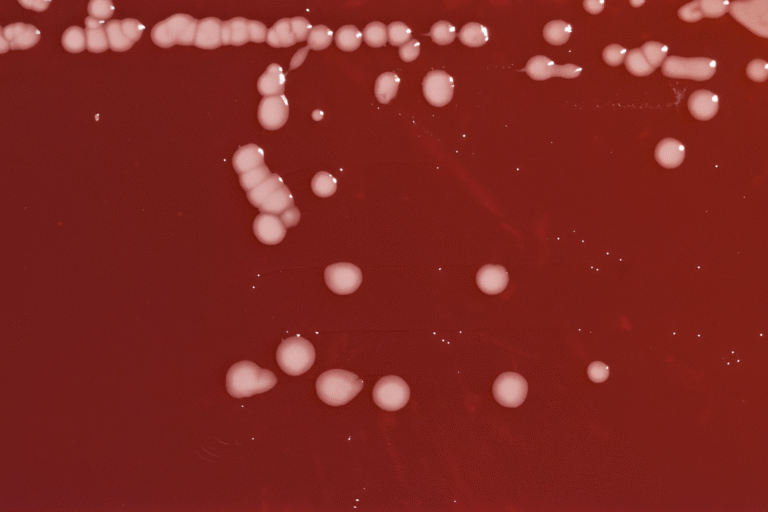
(A) Top and (B) bottom views of plates showing interactions between the phytopathogens Macrophomina sp. (M.sp), N. rosae (N.r), Fusarium sp. (F.sp), and Fusarium UG (F.UG) with the Trichoderma strains T. harzianum T9, T. harzianum M10, and T. atroviride IMI206040 (T.a). Phytopathogens are positioned on the right side, while Trichoderma strains are placed on the left. Photographs were taken after 96 hours of interaction.
(C) Close-up of the interaction zone between M.sp or N.r and the Trichoderma strains T9 and M10.
(D) Average colony area of M.sp and (E) N.r in interaction with different Trichoderma strains. The control represents the average colony size of each pathogen growing alone.
Asterisks indicate statistical differences according to the Tukey-HSD test at a significance level of α < 0.05, with p < 0.05. n = 3.

(A) Top view of plates showing interactions between T. harzianum T9, T. harzianum M10, and T. atroviride IMI206040 (T.a) with Macrophomina sp. (M.sp) or (C) N. rosae (N.r), cultivated on PDA medium at pH 8.5. Phytopathogens are shown on the right side, while Trichoderma strains are on the left. Photographs were taken after 96 hours of interaction.
(B) Average colony area of M.sp and (D) N.r in interaction with different Trichoderma strains. The control represents the average colony size of each pathogen growing alone.
Asterisks indicate statistical differences according to the Tukey-HSD test at a significance level of α < 0.05, with p < 0.05. n = 3.

(A) K-mer spectra analysis using Jellyfish with k = 21 for k-mer counting, and GenomeScope 1.0 for model fitting to estimate genome size, heterozygosity, and repetitiveness.
(B) Assessment of assembly completeness using BUSCO (Benchmarking Universal Single-Copy Orthologs) with the fungi dataset.
(C) Prediction and quantification of low-complexity regions and repetitive elements present in the genome.

(A) Table summarizing genome annotation statistics.
(B) Completeness of the annotation evaluated using BUSCO (Benchmarking Universal Single-Copy Orthologs) with the fungi dataset, based on transcripts extracted with GFFread.
(C) Gene length distribution.

(A) Rooted phylogenetic tree of 30 fungal species generated using OrthoVenn3 and ROADIES. Fusarium oxysporum and Fusarium graminearum were used as outgroup species.
(B) Venn diagram showing unique and shared orthologous cluster families among five Trichoderma species, including the sequenced T. harzianum T9 strain, generated with OrthoVenn3.

(A) Boxplot showing the percentage of SNPs in T9 orthologous genes compared with T. harzianum strains Th6, M10, T22, Th0179, Th3844, and TR274, using T. atroviride IMI206040 as a control.
(B) Scatterplot displaying the number of SNPs across orthologous genes in the comparison between T9 and the M10 strain.
(C) Annotation of genes with the highest SNP percentages (>11%) in the comparison of T9 versus M10.

This prediction was performed using fungiSMASH (the fungal analysis version of AntiSMASH).
Ten cluster families were identified, with T1PKS, NRPS, and terpene being the three most abundant.

(A) Extracted ion chromatograms of (to be completed with the m/z values of each extracted ion chromatogram).
(B) Fragmentation spectra of the precursor ion with m/z = 1175.7762 [M+H]+ corresponding to the 11-amino-acid peptaibol.
(C) Fragmentation spectra of the precursor ion with m/z = 1189.7918 [M+H]+ corresponding to the 11-amino-acid peptaibol.
(D) Fragmentation spectra of the precursor ion with m/z = 1442.9344 [M+H]+ corresponding to the 14-amino-acid peptaibol.

(A) Phylogenetic reconstruction using CORASON with “gene-name” as the query gene and the T. harzianum NRPS cluster responsible for 11- and 14-residue peptaibols. Genes absent in the reference cluster are highlighted and color-coded based on BLAST analysis.
(B) Biosynthetic pathway of the 12- and 14-residue peptaibols. The residues skipped during the biosynthetic process, caused by NRPS functionality, are shown in blue.
- Sariah, M., Choo, C. W., Zakaria, H. & Norihan, M. S. Quantification and characterisation of Trichoderma spp. from different ecosystems. Mycopathologia 159, 113–117 (2005).
- Woo, S. L., Hermosa, R., Lorito, M. & Monte, E. Trichoderma: a multipurpose, plant-beneficial microorganism for eco-sustainable agriculture. Nat. Rev. Microbiol. 21, 312–326 (2023).
- Guzmán-Guzmán, P., Porras-Troncoso, M. D., Olmedo-Monfil, V. & Herrera-Estrella, A. Trichoderma species: versatile plant symbionts. Phytopathology 109, 6–16 (2019).
- Tyśkiewicz, R., Nowak, A., Ozimek, E. & Jaroszuk-Ściseł, J. Trichoderma: the current status of its application in agriculture for the biocontrol of fungal phytopathogens and stimulation of plant growth. Int. J. Mol. Sci. 23, 2329 (2022).
- Yao, X., Guo, H., Zhang, K., Zhao, M., Ruan, J. & Chen, J. Trichoderma and its role in biological control of plant fungal and nematode disease. Front. Microbiol. 14, 1160551 (2023).
- Guo, R., Wang, Z., Huang, Y., Fan, H. & Liu, Z. Biocontrol potential of saline- or alkaline-tolerant Trichoderma asperellum mutants against three pathogenic fungi under saline or alkaline stress conditions. Braz. J. Microbiol. 49, 236–245 (2018).
- Bueno de Mesquita, C. P. et al. Microbial ecology and site characteristics underlie differences in salinity–methane relationships in coastal wetlands. J. Geophys. Res. Biogeosci. 129, e2024JG008133 (2024).
- Cabral-Miramontes, J. P., Olmedo-Monfil, V., Lara-Banda, M., Zúñiga-Romo, E. R. & Aréchiga-Carvajal, E. T. Promotion of plant growth in arid zones by selected Trichoderma spp. strains with adaptation plasticity to alkaline pH. Biology (Basel) 11, 1206 (2022). https://doi.org/10.3390/biology11081206
- Enriquez-Felix, E. E. et al. Argonaute and Dicer are essential for communication between Trichoderma atroviride and fungal hosts during mycoparasitism. Microbiol. Spectr. 12, e0316523 (2024). https://doi.org/10.1128/spectrum.03165-23
- Özkale, E., Yörük, E., Budak, M. & Korkmaz, E. M. Trichoderma atroviride suppresses Fusarium graminearum by altering primary and secondary metabolite biosynthesis profiling. Plant Pathol. 72, 1428–1441 (2023).
- Rebollar-Alviter, A., Silva-Rojas, H. V., Fuentes-Aragón, D., Acosta-González, U., Martínez-Ruiz, M. & Parra-Robles, B. E. An emerging strawberry fungal disease associated with root rot, crown rot and leaf spot caused by Neopestalotiopsis rosae in Mexico. Plant Dis. 104, 2054–2059 (2020).
- Chaverri, P., Branco-Rocha, F., Jaklitsch, W., Gazis, R., Degenkolb, T. & Samuels, G. J. Systematics of the Trichoderma harzianum species complex and the re-identification of commercial biocontrol strains. Mycologia 107, 558–590 (2015).
- Al-Salihi, S. A. & Alberti, F. Genomic based analysis of the biocontrol species Trichoderma harzianum: a model resource of structurally diverse pharmaceuticals and biopesticides. J. Fungi 9, 895 (2023).
- Sambrook, J. & Russell, D. W. Purification of nucleic acids by extraction with phenol: chloroform. Cold Spring Harb. Protoc. 2006, pdb-prot4455 (2006).
- White, T. J., Bruns, T., Lee, S. J. W. T. & Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications 315–322 (Academic Press, 1990).
- Swift, M. L. GraphPad Prism, data analysis, and scientific graphing. J. Chem. Inf. Comput. Sci. 37, 411–412 (1997).
- Andrews, S. FastQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
- Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
- Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
- Astashyn, A. et al. Rapid and sensitive detection of genome contamination at scale with FCS-GX. Genome Biol. 25, 60 (2024). https://doi.org/10.1186/s13059-024-03198-7
- Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013). https://doi.org/10.1093/bioinformatics/btt086
- Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
- Baril, T., Galbraith, J. G. & Hayward, A. Earl Grey: a fully automated user-friendly transposable element annotation and analysis pipeline. Mol. Biol. Evol. 41, msae068 (2024). https://doi.org/10.1093/molbev/msae068
- Hubley, R. et al. The Dfam database of repetitive DNA families. Nucleic Acids Res. 44, D81–D89 (2016).
- Bruna, T., Hoff, K. J., Lomsadze, A., Stanke, M. & Borodovsky, M. BRAKER2: automatic eukaryotic genome annotation with GeneMark-EP+ and AUGUSTUS supported by a protein database. NAR Genom. Bioinform. 3, lqaa108 (2021). https://doi.org/10.1093/nargab/lqaa108
- Kuznetsov, D. et al. OrthoDB v11: annotation of orthologs in the widest sampling of organismal diversity. Nucleic Acids Res. 51, D445–D451 (2023). https://doi.org/10.1093/nar/gkac998
- Tang, H. et al. JCVI: a versatile toolkit for comparative genomics analysis. Imeta 3, e211 (2024).
- Huerta-Cepas, J., Forslund, K., Coelho, L. P., Szklarczyk, D., Jensen, L. J., von Mering, C. & Bork, P. Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Mol. Biol. Evol. 34, 2115–2122 (2017). https://doi.org/10.1093/molbev/msx148
- Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014). https://doi.org/10.1093/bioinformatics/btu031
- Zheng, J., Ge, Q., Yan, Y., Zhang, X., Huang, L. & Yin, Y. dbCAN3: automated carbohydrate-active enzyme and substrate annotation. Nucleic Acids Res. 51, W115–W121 (2023).
- Cuomo, C. A. et al. The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science 317, 1400–1402 (2007). https://doi.org/10.1126/science.1143708
- Ma, L. J. et al. Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 464, 367–373 (2010). https://doi.org/10.1038/nature08850
- He, Y. & Zhu, H. Whole genome sequencing and analysis of the weed pathogen Trichoderma polysporum HZ-31. Sci. Rep. 14, 15228 (2024). https://doi.org/10.1038/s41598-024-66041-w
- Studholme, D. J. et al. Investigating the beneficial traits of Trichoderma hamatum GD12 for sustainable agriculture—insights from genomics. Front. Plant Sci. 4, 258 (2013). https://doi.org/10.3389/fpls.2013.00258
- Davolos, D. et al. A genomic and transcriptomic study on the DDT-resistant Trichoderma hamatum FBL 587: first genetic data into mycoremediation strategies for DDT-polluted sites. Microorganisms 9, 1680 (2021). https://doi.org/10.3390/microorganisms9081680
- Li, L., Zeng, X., Chen, J., Tian, J., Huang, J. & Su, S. Genome sequence of the fungus Trichoderma asperellum SM-12F1 revealing candidate functions of growth promotion, biocontrol, and bioremediation. PhytoFrontiers 1, 171–173 (2021). https://doi.org/10.1094/PHYTOFR-12-20-0052-A
- Li, W. C. et al. Complete genome sequences and genome-wide characterization of Trichoderma biocontrol agents provide new insights into their evolution and variation in genome organization, sexual development, and fungal–plant interactions. Microbiol. Spectr. 9, e0066321 (2021). https://doi.org/10.1128/Spectrum.00663-21
- Baroncelli, R., Zapparata, A., Piaggeschi, G., Sarrocco, S. & Vannacci, G. Draft whole-genome sequence of Trichoderma gamsii T6085, a promising biocontrol agent of Fusarium head blight on wheat. Genome Announc. 4, e01747-15 (2016). https://doi.org/10.1128/genomeA.01747-15
- Kubicek, C. P. et al. Comparative genome sequence analysis underscores mycoparasitism as the ancestral lifestyle of Trichoderma. Genome Biol. 12, R40 (2011). https://doi.org/10.1186/gb-2011-12-4-r40
- Druzhinina, I. S. et al. Massive lateral transfer of genes encoding plant cell wall-degrading enzymes to the mycoparasitic fungus Trichoderma from its plant-associated hosts. PLoS Genet. 14, e1007322 (2018). https://doi.org/10.1371/journal.pgen.1007322
- Kubicek, C. P. et al. Evolution and comparative genomics of the most common Trichoderma species. BMC Genomics 20, 485 (2019). https://doi.org/10.1186/s12864-019-5680-7
- Steindorff, A. S. et al. Identification of mycoparasitism-related genes against the phytopathogen Sclerotinia sclerotiorum through transcriptome and expression profile analysis in Trichoderma harzianum. BMC Genomics 15, 204 (2014). https://doi.org/10.1186/1471-2164-15-204
- Yang, D. et al. Genome sequence and annotation of Trichoderma parareesei, the ancestor of the cellulase producer Trichoderma reesei. Genome Announc. 3, e00885-15 (2015). https://doi.org/10.1128/genomeA.00885-15
- Li, W. C. et al. Trichoderma reesei complete genome sequence, repeat-induced point mutation, and partitioning of CAZyme gene clusters. Biotechnol. Biofuels 10, 170 (2017). https://doi.org/10.1186/s13068-017-0825-x
- Suazo Tejada, A. K. et al. Genome sequencing and de novo assembly of Trichoderma longibrachiatum isolate collected from Florida agricultural soils. Microbiol. Resour. Announc. 13, e00906-23 (2024). https://doi.org/10.1128/mra.00906-23
- Rosolen, R. R. et al. Whole-genome sequencing and comparative genomic analysis of potential biotechnological strains of Trichoderma harzianum, Trichoderma atroviride, and Trichoderma reesei. Mol. Genet. Genomics 298, 735–754 (2023). https://doi.org/10.1007/s00438-023-02013-5
- Sun, J., Lu, F., Luo, Y., Bie, L., Xu, L. & Wang, Y. OrthoVenn3: an integrated platform for exploring and visualizing orthologous data across genomes. Nucleic Acids Res. 51, W397–W403 (2023).
- Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004). https://doi.org/10.1093/nar/gkh340
- Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009). https://doi.org/10.1093/bioinformatics/btp348
- Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5, e9490 (2010).
- Gupta, A., Mirarab, S. & Turakhia, Y. Accurate, scalable, and fully automated inference of species trees from raw genome assemblies using ROADIES. Proc. Natl Acad. Sci. USA 122, e2500553122 (2025). https://doi.org/10.1073/pnas.2500553122
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv (2013). https://doi.org/10.48550/arXiv.1303.3997
- Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009). https://doi.org/10.1093/bioinformatics/btp352
- Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011). https://doi.org/10.1093/bioinformatics/btr330
- Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019). https://doi.org/10.1186/s13059-019-1832-y
- Heuckeroth, S. et al. Reproducible mass spectrometry data processing and compound annotation in MZmine 3. Nat. Protoc. 19, 2597–2641 (2024).
- Navarro-Muñoz, J. C. et al. A computational framework to explore large-scale biosynthetic diversity. Nat. Chem. Biol. 16, 60–68 (2020).
- Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
- Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204 (2017).
- Baroncelli, R. et al. Draft whole-genome sequence of the biocontrol agent Trichoderma harzianum T6776. Genome Announc. 3, e00647-15 (2015). https://doi.org/10.1128/genomeA.00647-15
- Fanelli, F., Liuzzi, V. C., Logrieco, A. F. & et al. Genomic characterization of Trichoderma atrobrunneum (T. harzianum species complex) ITEM 908: insight into the genetic endowment of a multi-target biocontrol strain. BMC Genomics 19, 662 (2018). https://doi.org/10.1186/s12864-018-5049-3
- Beijen, E. P. W. & Ohm, R. A. Genome annotations for the ascomycete fungi Trichoderma harzianum, Trichoderma aggressivum and Purpureocillium lilacinum. Microbiol. Resour. Announc. 13, e01153-23 (2024). https://doi.org/10.1128/mra.01153-23
- Pertea, G. & Pertea, M. GFF utilities: GffRead and GffCompare. F1000Research 9, ISCB-Comm (2020).
- Dixon, D. P., Skipsey, M. & Edwards, R. Roles for glutathione transferases in plant secondary metabolism. Phytochemistry 71, 338–350 (2010).
- Baker, S. E., Perrone, G., Richardson, N. M., Gallo, A. & Kubicek, C. P. Phylogenomic analysis of polyketide synthase-encoding genes in Trichoderma. Microbiology 158, 147–154 (2012).
- Bumpus, S. B., Magarvey, N. A., Kelleher, N. L., Walsh, C. T. & Calderone, C. T. Polyunsaturated fatty-acid-like trans-enoyl reductases utilized in polyketide biosynthesis. J. Am. Chem. Soc. 130, 11614–11616 (2008).
- Yao, T., Wang, X. & Chen, F. The role of enoyl reductase in the Monacolin K biosynthesis pathway in Monascus spp. J. Fungi 11, 199 (2025).
- Moummou, H., Kallberg, Y., Tonfack, L. B., Persson, B. & van Der Rest, B. The plant short-chain dehydrogenase (SDR) superfamily: genome-wide inventory and diversification patterns. BMC Plant Biol. 12, 219 (2012).
- Mukherjee, P. K. et al. Two classes of new peptaibols are synthesized by a single non-ribosomal peptide synthetase of Trichoderma virens. J. Biol. Chem. 286, 4544–4554 (2011).
- Nandini, B., Puttaswamy, H., Saini, R. K., Prakash, H. S. & Geetha, N. Trichovariability in rhizosphere soil samples and their biocontrol potential against downy mildew pathogen in pearl millet. Sci. Rep. 11, 9517 (2021). https://doi.org/10.1038/s41598-021-89061-2
- Xue, M. et al. Screening and identification of Trichoderma strains isolated from natural habitats in China with potential agricultural applications. Biomed. Res. Int. 2021, 7913950 (2021). https://doi.org/10.1155/2021/7913950
- Mukherjee, P. K., Mendoza-Mendoza, A., Zeilinger, S. & Horwitz, B. A. Mycoparasitism as a mechanism of Trichoderma-mediated suppression of plant diseases. Fungal Biol. Rev. 38, 15–33 (2022). https://doi.org/10.1016/j.fbr.2021.11.004
- Timmusk, S., Nevo, E., Ayele, F., Noe, S. & Niinemets, Ü. Fighting Fusarium pathogens in the era of climate change: a conceptual approach. Pathogens 9, 419 (2020).
- Singh, N. K., Tralamazza, S. M., Abraham, L. N., Glauser, G. & Croll, D. Genome-wide association mapping reveals genes underlying population-level metabolome diversity in a fungal crop pathogen. BMC Biol. 20, 224 (2022).
- Mukherjee, P. K., Horwitz, B. A. & Kenerley, C. M. Secondary metabolism in Trichoderma – a genomic perspective. Microbiology 158, 35–45 (2012).